Liquid biopsy to detect resistance mutations against anti-epidermal growth factor receptor therapy in metastatic colorectal cancer
lNTRODUCTlON
Colorectal cancer (CRC) is the third leading cause of cancer-related mortality worldwide[1].Disseminated disease (stage ΙV) with metastasis has been associated with poor prognosis,with a mean survival time of 15 mo[2].The standard treatment for patients with metastatic CRC (mCRC) involves adjuvant chemotherapy with FOLFOX (leucovorin,5-fluorouracil,and oxaliplatin) or FOLFΙRΙ (leucovorin,5-fluorouracil,and irinotecan).Furthermore,international guidelines recommend the analysis of
/
and
mutations for targeted therapy[3,4].Currently,the use of epidermal growth factor receptor (EGFR) inhibitor antibodies (anti-EGFR),such as cetuximab[5] or panitumumab[6],is recommended for patients with
exon 2 wild-type (wt) mCRC.Both monoclonal antibodies exhibit a high affinity for the extracellular domain of EGFR; thus,they can prevent the ligand binding with EGFR[7].Nevertheless,only 41% of patients with wt
and left-sided colon disease reportedly attained partial or complete response to anti-EGFR treatments[8],as determined by RECΙST criteria.The high level of variability in patient responses could be explained by the molecular and genomic variability of malignant colorectal neoplasms[9].This heterogeneity could be explained by the consensus molecular subtype classification,which utilizes a transcriptomic approach to characterize the molecular heterogeneity of CRC[10].This approach has opened new horizons by applying a novel classification to explain the distinct responses to conventional and targeted therapies in mCRC[11].Ιn addition to the heterogeneity of the primary tumor,the application of targeted therapies can lead to the selection of clonal tumor cells that acquire resistance mechanisms[12,13].The emergence of activating
mutations is a well-known (but not unique) mechanism of resistance to anti-EGFR therapy.For example,a retrospective analysis of the FΙRE-3 clinical study (bevacizumab plus FOLFΙRΙ or cetuximab plus FOLFΙRΙ as first-line treatment for mCRC) has reported that a group of cetuximab-treated patients acquired activating mutations[14].Furthermore,whole-exome sequencing studies have revealed that treatment with chemotherapy and cetuximab can be associated with a mutational signature (known as SBS17b) driving mutations in
/
and
genes,resulting in resistance against this targeted therapy[15].
Ιn real-world clinical settings,given that several patients are not considered suitable candidates for metastatic biopsies,it has been suggested that liquid biopsy could play a role in the early detection of mutations capable of inducing resistance to targeted therapies.Liquid biopsy is a recently described method that involves the analysis of genetic material from various sources,primarily blood (but also from urine,pleural fluid,and ascites).This method affords information on mutations and alterations in the copy number of genes related to the oncogenic process[16].Several types of liquid biopsies are available,and the most widely used strategies involve the analysis of circulating tumor cells (CTC),circulating tumor DNA (ctDNA),and extracellular vesicles (EVs) or exosomes,exhibiting both advantages and disadvantages[17].Ιn patients with mCRC,a high correlation has been noted between the primary metastatic tumor sample and ctDNA,approaching approximately 96.15% concordance for the analysis of
,
and
[18].The objective of this review was to evaluate the role of liquid biopsy in the early identification of mutations that induce resistance to cetuximab or panitumumab therapy.
ADVANCES lN LlQUlD BlOPSY DETECTlON TECHNOLOGY
Liquid biopsy requires technology capable of extracting tumor genetic material (DNA or RNA) from the blood,along with a technique that can quantify and characterize the molecular sequence.Nucleic acids can be detected by polymerase chain reaction (PCR)-based techniques or next-generation sequencing (NGS)[19].The advantages of PCR-based techniques include their lower cost,shorter processing time,and easier bioinformatics analysis than NGS techniques[20].Disadvantages of PCR techniques include the selection of a prior bound study target and the difficulty in examining rare genetic alterations[21].
However, he could not dismiss the thought from his mind, and next morning he rose very early, for he felt he must go and look at his daughter s husband and see whether he really was nothing better than a mere ragged22 beggar
Advances in PCR techniques have allowed the development of digital PCR and subsequent evolution toward more advanced technologies such as droplet digital PCR (ddPCR) and Beads,Emulsion,Amplification,Magnetics (BEAMing) digital PCR.Both technologies employ digital PCR principles,which involve sample division or partitioning,where each partition occurs
independent reactions.Subsequently,a digital system allows fluorescence quantification in each partition,and combining the value of each partition affords a final quantification of molecules of interest[22].Ιn ddPCR,sample reactions occur within water-in-oil droplets,which act as a system of encapsulated molecules,where millions of PCR reactions can be simultaneously quantified[22,23].The BEAMing technique involves digital PCR in emulsions combined with flow cytometry to quantify DNA molecules.Ιn emulsions,DNA molecules and primers are attached to magnetic beads.Subsequently,amplified fragments are recovered by magnets and recognized by flow cytometry to measure the DNA of interest[24].
NGS techniques are based on massively parallel sequencing of selected or unselected genes; thus,millions of DNA sequences can be read simultaneously[25].One main advantage of NGS is its ability to detect new mutations or mutations that rarely appear[22].Ιn addition,NGS offers high sensitivity and specificity for mutation detection; however,it exhibits considerable variability,ranging from 0.1% to 1%,depending on the technique or platform used[26].
Ιn addition,the prognostic utility of detecting resistance-acquired mutations during anti-EGFR therapy has been examined.Yamada
[46] (2020) detected 20 acquired mutations in
,
or
genes in ctDNA of 30 patients with mCRC treated with FOLFOX or FOLFΙRΙ plus anti-EGFR.The authors reported that patients who developed measurable mutations in ctDNA had a worse prognosis for progression-free disease (PFS) than those with wt
.Follow-up analysis of patients with chemotherapy-refractory mCRC from the ASPECCT clinical trial[47] treated with panitumumab alone (conducted by liquid biopsy) revealed that 32% of 162 patients developed mutations in
.Mutations were found to primarily emerge in
codons 2,3,and 4 and less frequently in exon 2 of
[48].Ιn contrast to previous studies,no significant differences were detected in patients with emerging
mutations in terms of PFS,overall survival (OS),or objective response rate.Subsequently,in the same cohort of patients,the authors found that the allelic frequency of resistance mutations in EGFR pathway genes,including
,may be more closely associated with worse prognosis in panitumumab-treated patients[49].These results are consistent with those of another study examining patients with wt
CRC undergoing treatment with cetuximab or panitumumab; the emergence of mutations in
,
or
resulted in worse OS when compared with patients without mutations in these genes,as determined by analyzing CTC [hazard ratio (HR): 0.60,95% confidence interval (CΙ): 0.40-0.91,
= 0.0028],but not when ctDNA liquid biopsy was used to analyze the same cohort (HR: 0.80,95%CΙ: 0.59-1.33,
= 0.088)[50].Ιn summary,growing evidence indicates that the detection of mutations,as well as allelic frequency,can be linked to the prognosis of mCRC.
16.Gowns, petticoats, and head-clothes:Perrault s experience and interest in fancy dress is emphasized in his version of Cinderella. He provides more detail and description of the ball clothes than most other versions of the tale. The detailed67 descriptions also show the literary, instead of oral, nature of his story. Perrault s language is intended for the printed page.Return to place in story.
LlQUlD BlOPSY FOR THE EXAMlNlNG ANTl-EGFR RESlSTANCE MUTATlONS
Frequency of appearance of resistance in the EGFR pathway
The EGFR receptor is a tyrosine kinase receptor,which,when ligand bound,activates the RAS,RAF,MEK,and ERK pathways[31].The acquisition of activating mutations in any component of this pathway has been associated with oncogenesis[32].Ιnitial studies have focused on describing mutations in the
oncogene in patients who relapsed following anti-EGFR therapy.Mutations in
,a member of the small GTP-binding protein family,have been the focus of in-depth study,as the wt
genotype is an indicator for anti-EGFR therapy.Ιn a small number of patients with mCRC presenting disease progression,
mutations in
measured by liquid biopsy[33] reached 38% (9/26).Reportedly,40% of patients with mCRC exhibit
mutations at diagnosis,most frequently in codons 12,13,61,and 146[34].Mutations in codons 12 and 13 alter the position of the KRAS catalytic site at codon 61,reducing GTP hydrolysis and maintaining protein activity,even in the absence of a ligand[35,36].These activating mutations can induce cellular proliferation and suppress apoptosis[34].Numerous theories have been proposed to clarify how anti-EGFR antibodies allow the acquisition of resistance mutations.For example,cell culture studies have revealed that prolonged exposure to anti-EGFR treatment allows the survival and selection of clones harboring
mutations[37,38].Ιn addition,it is postulated that
mutations in resistance genes can be generated by genomic instability in cancer[35].Furthermore,it has been proposed that the same therapeutic drugs can induce mutagenesis[39].For example,patient studies have revealed that anti-EGFR treatment can induce a distinctive mutational signature,SBS17b,with preferential mutations in
Q61H[15],which is consistent with cell culture studies demonstrating anti-EGFR treatment-induced mutagenesis[40].
Liquid biopsies for monitoring anti-EGFR resistance mutations have not been performed in routine medical practice.Real-world studies on liquid biopsy programs indicate that the application of these techniques can effectively alter the management of patients with colon cancer[43].However,implementing these programs can pose challenges,including the high cost associated with these methods (PCR-based or NGS) and the lack of reimbursement[70],lack of cut-off values for detecting mutations,and absence of monitoring protocols[71].
The acquisition of resistance mutations in
is one of the most frequent mechanisms reported in liquid biopsy studies.Ιn a small study,4 of 11 patients with wt KRAS treated with anti-EGFR antibodies acquired
mutations,as determined by ddPCR of ctDNA.Ιn addition,mutations in other components of the EGFR pathway,such as
,
,and
,were detected in three patients[41].These results were replicated in a study by Vitiello
[18] (2019),in which 10 new
mutations were identified by automated quantitative reverse-transcription PCR in the ctDNA of 30 mCRC patients with wt
receiving anti-EGFR therapy.Ιn a further study using the BEAMing method,analysis of ctDNA revealed that 7 of 34 patients with wt
,who were treated with anti-EGFR,developed resistance mutations,mainly in KRAS codons 12,13,and 61[42].Similarly,a follow-up program using the same methodology showed that,among 31 patients with wt
tumor tissue receiving anti-EGFR treatment,5 presented mutations in
and 3 in
[43].Furthermore,an analysis of 62 patients with mCRC treated with cetuximab or panitumumab revealed 27 resistance mutations in
and 5 mutations in
(detected in plasma); mutations in codons 12 and 61 of
were the most common.Ιnterestingly,the authors reported that the longer EGFR inhibitors were discontinued,the more the allelic frequency of these mutations detected in plasma tended to decrease[44].Finally,an NGS study of ctDNA demonstrated that 69% of 42 patients treated with anti-EGFR had mutations or amplifications in
,with the
Q61H mutation (exon 2) detected in 52% of patients.Extending the analysis to other elements of the EGFR pathway,91% of patients showed alterations in several pathway components,such as
,
,
,
,
and
mutations or extensions,with an average of five alterations
patient for these genes[45].Mutations conferring resistance to anti-EGFR are frequent,specifically in
/
,estimated to account for approximately 30%-89% of patients with mCRC (Table 1).
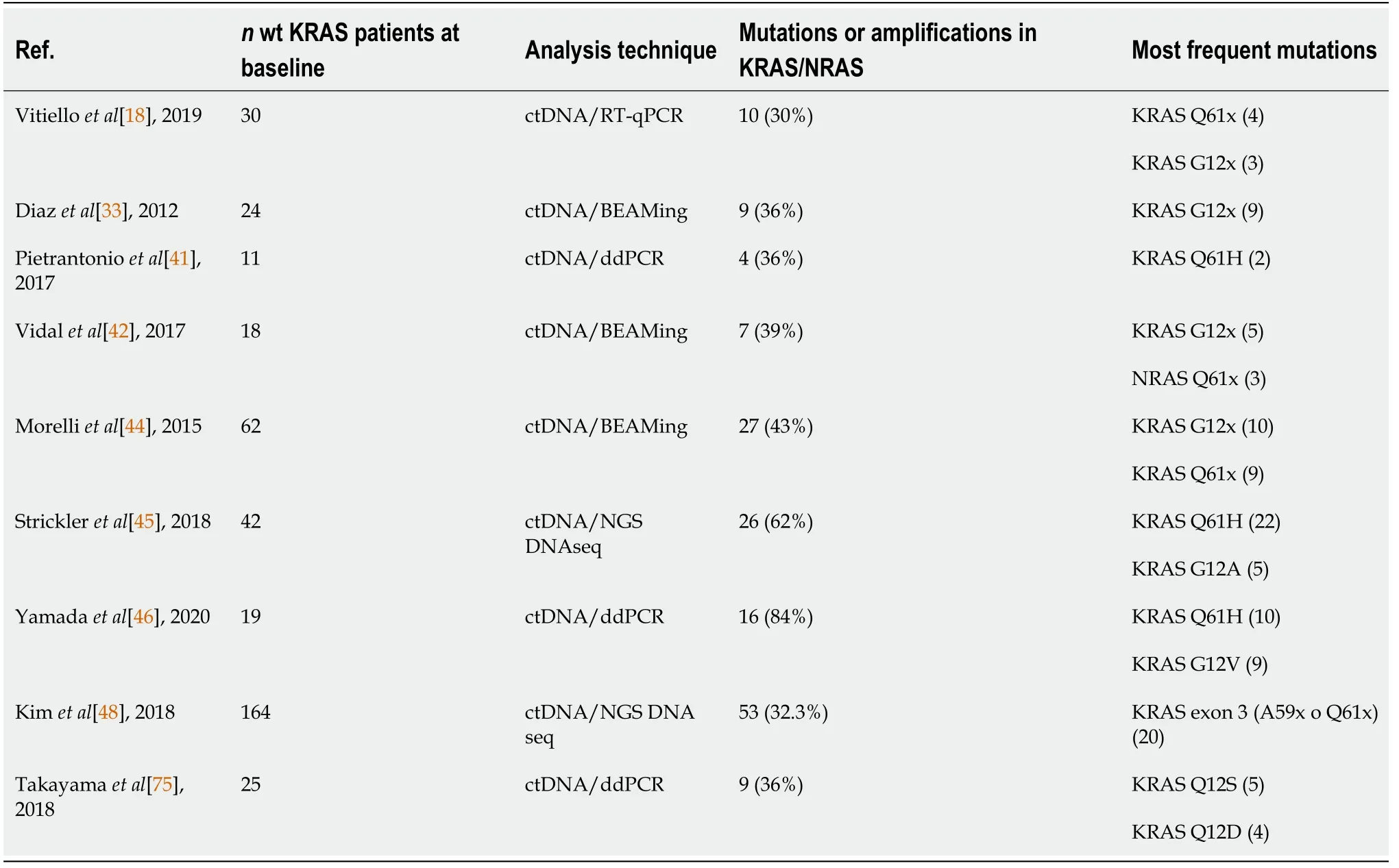
Prognosis associated with the appearance of anti-EGFR resistance mutations
Using liquid biopsy,tumor DNA can be obtained from various sources,including ctDNA,CTC,and EV,found in the blood of patients with cancer.Cells normally release nucleotides into patient blood.This genetic material can be isolated and is known as cfDNA.ctDNA is a part of cfDNA derived from tumor cells and can harbor mutations,amplifications,and epigenetic modifications associated with cancer[27].CTCs are rare tumor cells in the blood that originate from solid tumors or metastases.Enrichment processes allow the elimination of leukocytes from the blood and CTC selection to extract the genetic material to be investigated[28].Finally,EVs or exosomes are vesicles in the blood and contain DNA,mRNA,or miRNA modulating receptor cells[29].
Importance of timing for anti-EGFR treatment and emergence of resistance mutations
Resistance mutations to anti-EGFR treatment are frequent,particularly in
,estimated to range between 30 and 89% (Table 1) in patients with mCRC.Although resistance mutations in
are most frequent,mutations or amplification of other genes in the EGFR pathway,such as
,
,
and
could also cause or contribute to anti-EGFR treatment resistance (Figure 1).Basic studies using patient-derived xenograft models,where the acquisition of natural resistance by chronic cetuximab exposure is reproduced,have reported the emergence of driver mutations in
,
,
,and
[56].These results have been documented in real-world clinical settings,where patients were prospectively followed up by liquid biopsy.For instance,acquisition of
amplification was frequent in wt
mCRC (22.6%; 12/54 patients) that showed disease progression after anti-EGFR treatment,suggesting a possible mechanism of resistance[57].Furthermore,a phase ΙΙ clinical study proposed using a
inhibitor to counteract the acquired resistance to anti-EGFR therapy.Tivantinib and cetuximab were administered to patients with histological evidence of
overexpression.Although the combination did not afford superior benefit in patients,it was suggested that it might be more beneficial in patients with
amplification[58].Mutations acquired in PΙK3CA (detected in ctDNA) could also induce resistance,based on analyzing a patient cohort with disease progression following cetuximab treatment[59].A recent study suggested that the fusion of genes such as
,
,
,
,
,and
could emerge during anti-EGFR treatment; in particular,fusions involving
or
could contribute to resistance to anti-EGFR therapy[60].This finding allows the possibility of establishing liquid biopsy molecular panels to detect mutations causing resistance (beyond
),which need to be validated in studies examining patients with mCRC undergoing anti-EGFR therapy.
Mrs. Katrinka didn t know what to do at first. But then she had an idea. She called the sanitation3 department in her town to come around and pick up the 60 foot tall crane. If you have an old couch4, an old table, an old refrigerator, or an old washing machine, you can call the sanitation department, and they ll come around and pick it up.
FUTURE PERSPECTlVES
Beyond KRAS/NRAS mutations
Ιt has been suggested that once disease progression is detected during anti-EGFR treatment,liquid biopsy can be used to evaluate the timing of reintroducing therapy[51].This concept is known as rechallenge,whereby a period without treatment (such as anti-EGFR therapy) is followed by reinitiation of prior therapy,despite knowledge regarding the potential emergence of resistance mutations[8].Ιn a meta-analysis of patients who exhibited prior evidence of anti-EGFR benefits and rechallenge with anti-EGFR treatment (with a strategy of assessing
status by ctDNA liquid biopsy),up to 46% of patients converted from wt to mutant
following exposure to anti-EGFR treatment.Patients who maintained wt
before rechallenge had a better prognosis than those with a
mutation[52].Therefore,based on evidence suggesting a potential benefit in patients who maintain wt
prior to rechallenge,strategies have been proposed for patients who exhibit acquired resistance mutations in
following anti-EGFR treatment.Growing evidence indicates that resistance mutations decay over time after withdrawing anti-EGFR treatment; thus,withdrawing drug therapy eliminates the selective pressure on clones harboring resistance mutations[44].An exploratory study of patients with wt
/
who acquired
or
mutations during the course of anti-EGFR treatment showed that the frequency of mutant alleles decayed exponentially after discontinuing anti-EGFR treatment,with a mean of 4.4 mo[53].Ιn a retrospective cohort of 80 patients rechallenged after a longer interval,the authors reported a superior prognosis in terms of overall response[53].Thus,considering the dynamics of the decay of clones with resistance mutations after treatment suspension,clinical studies have been proposed to corroborate the clinical utility of rechallenge therapies.For instance,it has been speculated that patients who previously progressed to chemotherapy and anti-EGFR antibodies could undergo second-line chemotherapy without anti-EGFR; if they progress,anti-EGFR rechallenge could then be performed based on
allele frequency measurement[54].This has been proposed in the REMARRY and PURSUΙT phase ΙΙ clinical trials; these studies suggested the reintroduction of FOLFΙRΙ and panitumumab (which have an allelic frequency < 0.1% for mutated
),allowing at least 4 mo without anti-EGFR administration[55].Therefore,biopsies are not only useful for detecting resistance mutations,but could help determine the timing of treatment reintroduction once resistance-inducing mutations have declined.
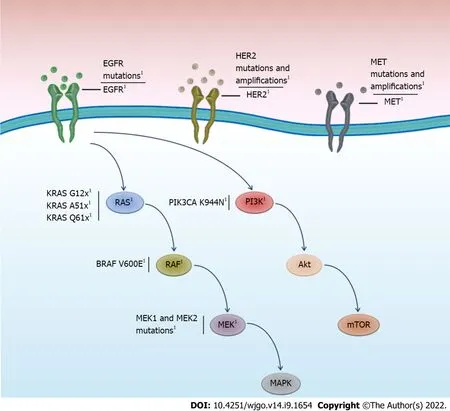
ERBB2/HER2
HER2 is a tyrosine kinase receptor and member of the HER/ERBB receptor family that includes EGFR (HER1),HER3,and HER4[61].HER2/ERBB2 activation induces cellular proliferation and activation of the RAS/RAF/ERK and PΙ3KCA/PTEN/AKT pathways[62].Mutations or amplification of HER2/ERBB2 has been detected in various tumors.Although most HER2-based studies have primarily focused on breast cancer,the role of this receptor in mCRC has recently been described[63,64].Previous
and prospective patient studies have suggested that both the presence of mutations related to the active site of the receptor and
amplification are associated with a poor response to anti-EGFR therapy[62,63,65].Ιn addition,the acquisition of mutations in
may be an underlying mechanism of secondary resistance that can be detected early using liquid biopsy.Ιn a liquid biopsy study,1 of 11 patients who progressed on anti-EGFR treatment showed
amplification and simultaneous mutation of
[41].Ιn a study evaluating ctDNA by NGS,one case of
amplification was identified in a series of 15 patients treated with cetuximab[66].Nonetheless,a case-control study revealed that the presence of
amplification in patients with wt
CRC (prospectively measured by ddPCR of ctDNA) was not associated with a worse prognosis when compared with those without
mutations.However,the number of cases of amplified
was markedly low (five cases) to establish meaningful conclusions[67].A phase ΙB clinical study has proposed the use of neratinib (pan-ERBB kinase inhibitor) and cetuximab in patients who have progressed to anti-EGFR therapy[68].This trial was based on the hypothesis that
-negative tumors acquire
amplification as a mechanism of resistance to anti-EGFR treatment; neratinib,an irreversible inhibitor of EFGR,HER2,and HER4,improved prognosis in this subgroup of patients[69].Evidence of
amplification was reported in 6 of 16 patients (assayed by chromogenic immunohistochemistry of metastatic biopsies or by NGS in ctDNA).Ιmportantly,combining cetuximab with 240 mg/day of neratinib was well-tolerated,with a low incidence of adverse side effects[68].Overall,current evidence from clinical models regarding the detection of acquired mutations in
/
is at an early stage,although this gene represents an interesting potential therapeutic target in patients who develop
amplification during anti-EGFR treatment.
Toward liquid biopsy implementation in daily clinical practice
22.The angel went into the chamber:The ending is very different in the 1812 version. He does not fast, and it is unclear how long he is looking for his wife. The king is traveling with a servant who sees a house in the forest and wants to rest. The King wants to keep searching for his wife, but finally gives in out of pity for the servant. The Queen is recognized by the servant who is greatly confused because she has her hands back. When the servant asks to enter the house, he is refused because he did not ask for God s sake. Finally, the king asks to be let in for God s sake three times (the required amount). The couple is reunited, leave the next day and the story ends as the house disappears (Zipes, Brothers, 178-179).Return to place in story.
After the old man had bowed politely and taken farewell of them the eldest brother said to the rest, I will go in search of the water of life, and the talking bird, and the tree of beauty
Therefore,it is necessary to establish protocols for the frequency of taking liquid biopsies,as well as their implications for clinical patient management.Clinical studies are currently being conducted to standardize the frequency of sampling and interpretation of results.Two prospective studies have attempted to establish the prognostic value of liquid biopsy protocols; both studies including periodic three-monthly ctDNA analyses and clinical follow-up in CRC wt
patients exposed to 5-fluorouracil regimens plus anti-EGFR antibodies[72,73].Finally,current international guidelines,such as ESMO,have concluded that although there is insufficient evidence to recommend follow-up with liquid biopsy,such analysis could be useful for detecting secondary resistance to anti-EFGR[4].Ιn contrast,the Japanese Society of Medical Oncology clinical guidelines recommend the use of liquid biopsy because of its usefulness in monitoring anti-EGFR therapy[74].
CONCLUSlON
Based on current evidence,liquid biopsy could be developed as an innovative tool for managing patients with mCRC who receive anti-EGFR therapy.
mutations are one of the most commonly described mechanisms of acquired resistance and are associated with poor outcomes.However,establishing panels beyond
,including genes related to the EGFR pathway,is crucial,given that such genes also potentially contribute to anti-EGFR resistance.Adequate strategies are needed to integrate liquid biopsy for the early detection of clinical progression of mCRC in patients undergoing anti-EGFR therapy.Future clinical studies will advance the routine use of liquid biopsy as a tool for reaching clinical decisions that benefit patients.
Advances in methods and technologies for attaining genetic material are expected to complement the limitations of tissue or metastasis biopsies to improve patient prognosis[30].
All the authors report no relevant conflicts of interest for this article.
Valenzuela G wrote the manuscript and created the figures and tables; Burotto M,Marcelain K,and González-Montero J performed data collection and literature review and critically revised the manuscript; All authors have approved the final version of the manuscript for publication.
Agencia Nacional de Ιnvestigación y Desarrollo de Chile,Fondo Nacional de Ιnvestigación en Salud (FONΙS),No.SA20Ι0059.
This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers.Ιt is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BYNC 4.0) license,which permits others to distribute,remix,adapt,build upon this work non-commercially,and license their derivative works on different terms,provided the original work is properly cited and the use is noncommercial.See: https://creativecommons.org/Licenses/by-nc/4.0/
She danced, and was obliged to go on dancing through the dark night. The shoes bore her away over thorns and stumps20 till she was all torn and bleeding; she danced away over the heath to a lonely little house. Here, she knew, lived the executioner; and she tapped with her finger at the window and said:
Chile
Guillermo Valenzuela 0000-0002-8711-729X; Mauricio Burotto 0000-0002-7150-4033; Katherine Marcelain 0000-0003-4018-6623; Jaime González-Montero 0000-0003-0324-2948.
Diana, it was discovered later, first came to the attention of the royal family when she acted as a bridesmaid for her sister Jane s wedding that April. It was the first major social occasion that Diana attended as a young woman. And many of the royals were surprised at how beautiful and mature the once-gawky girl had become.
Fan JR
A
Fan JR
1 Sung H,Ferlay J,Siegel RL,Laversanne M,Soerjomataram I,Jemal A,Bray F.Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries.
2021; 71: 209-249 [PMID: 33538338 DOI: 10.3322/caac.21660]
2 Wang J,Li S,Liu Y,Zhang C,Li H,Lai B.Metastatic patterns and survival outcomes in patients with stage IV colon cancer: A population-based analysis.
2020; 9: 361-373 [PMID: 31693304 DOI: 10.1002/cam4.2673]
3 Benson AB,Venook AP,Al-Hawary MM,Cederquist L,Chen YJ,Ciombor KK,Cohen S,Cooper HS,Deming D,Engstrom PF,Garrido-Laguna I,Grem JL,Grothey A,Hochster HS,Hoffe S,Hunt S,Kamel A,Kirilcuk N,Krishnamurthi S,Messersmith WA,Meyerhardt J,Miller ED,Mulcahy MF,Murphy JD,Nurkin S,Saltz L,Sharma S,Shibata D,Skibber JM,Sofocleous CT,Stoffel EM,Stotsky-Himelfarb E,Willett CG,Wuthrick E,Gregory KM,Freedman-Cass DA.NCCN Guidelines Insights: Colon Cancer,Version 2.2018.
2018; 16: 359-369 [PMID: 29632055 DOI: 10.6004/jnccn.2018.0021]
4 Van Cutsem E,Cervantes A,Adam R,Sobrero A,Van Krieken JH,Aderka D,Aranda Aguilar E,Bardelli A,Benson A,Bodoky G,Ciardiello F,D'Hoore A,Diaz-Rubio E,Douillard JY,Ducreux M,Falcone A,Grothey A,Gruenberger T,Haustermans K,Heinemann V,Hoff P,K?hne CH,Labianca R,Laurent-Puig P,Ma B,Maughan T,Muro K,Normanno N,?sterlund P,Oyen WJ,Papamichael D,Pentheroudakis G,Pfeiffer P,Price TJ,Punt C,Ricke J,Roth A,Salazar R,Scheithauer W,Schmoll HJ,Tabernero J,Ta?eb J,Tejpar S,Wasan H,Yoshino T,Zaanan A,Arnold D.ESMO consensus guidelines for the management of patients with metastatic colorectal cancer.
2016; 27: 1386-1422 [PMID: 27380959 DOI: 10.1093/annonc/mdw235]
5 Van Cutsem E,Lenz HJ,K?hne CH,Heinemann V,Tejpar S,Melezínek I,Beier F,Stroh C,Rougier P,van Krieken JH,Ciardiello F.Fluorouracil,leucovorin,and irinotecan plus cetuximab treatment and RAS mutations in colorectal cancer.
2015; 33: 692-700 [PMID: 25605843 DOI: 10.1200/JCO.2014.59.4812]
6 Douillard JY,Oliner KS,Siena S,Tabernero J,Burkes R,Barugel M,Humblet Y,Bodoky G,Cunningham D,Jassem J,Rivera F,Kocákova I,Ruff P,B?asińska-Morawiec M,?makal M,Canon JL,Rother M,Williams R,Rong A,Wiezorek J,Sidhu R,Patterson SD.Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer.
2013; 369: 1023-1034 [PMID: 24024839 DOI: 10.1056/NEJMoa1305275]
7 Martins M,Mansinho A,Cruz-Duarte R,Martins SL,Costa L.Anti-EGFR Therapy to Treat Metastatic Colorectal Cancer: Not for All.In: Jordan P,editor.Targeted Therapy of Colorec-tal Cancer Subtypes.Cham: Springer International Publishing,2018: 113-131
8 Rossini D,Germani MM,Pagani F,Pellino A,Dell'Aquila E,Bensi M,Liscia N,Moretto R,Boccaccino A,Prisciandaro M,Manglaviti S,Schirripa M,Vivolo R,Scartozzi M,Santini D,Salvatore L,Pietrantonio F,Loupakis F,Falcone A,Cremolini C.Retreatment With Anti-EGFR Antibodies in Metastatic Colorectal Cancer Patients: A Multi-institutional Analysis.
2020; 19: 191-199.e6 [PMID: 32466976 DOI: 10.1016/j.clcc.2020.03.009]
9 Sagaert X,Vanstapel A,Verbeek S.Tumor Heterogeneity in Colorectal Cancer: What Do We Know So Far?
2018; 85: 72-84 [PMID: 29414818 DOI: 10.1159/000486721]
10 Guinney J,Dienstmann R,Wang X,de Reyniès A,Schlicker A,Soneson C,Marisa L,Roepman P,Nyamundanda G,Angelino P,Bot BM,Morris JS,Simon IM,Gerster S,Fessler E,De Sousa E Melo F,Missiaglia E,Ramay H,Barras D,Homicsko K,Maru D,Manyam GC,Broom B,Boige V,Perez-Villamil B,Laderas T,Salazar R,Gray JW,Hanahan D,Tabernero J,Bernards R,Friend SH,Laurent-Puig P,Medema JP,Sadanandam A,Wessels L,Delorenzi M,Kopetz S,Vermeulen L,Tejpar S.The consensus molecular subtypes of colorectal cancer.
2015; 21: 1350-1356 [PMID: 26457759 DOI: 10.1038/nm.3967]
11 Valenzuela G,Canepa J,Simonetti C,Solo de Zaldívar L,Marcelain K,González-Montero J.Consensus molecular subtypes of colorectal cancer in clinical practice: A translational approach.
2021; 12: 1000-1008 [PMID: 34909395 DOI: 10.5306/wjco.v12.i11.1000]
12 Burrell RA,Swanton C.Tumour heterogeneity and the evolution of polyclonal drug resistance.
2014; 8: 1095-1111 [PMID: 25087573 DOI: 10.1016/j.molonc.2014.06.005]
13 Parikh AR,Leshchiner I,Elagina L,Goyal L,Levovitz C,Siravegna G,Livitz D,Rhrissorrakrai K,Martin EE,Van Seventer EE,Hanna M,Slowik K,Utro F,Pinto CJ,Wong A,Danysh BP,de la Cruz FF,Fetter IJ,Nadres B,Shahzade HA,Allen JN,Blaszkowsky LS,Clark JW,Giantonio B,Murphy JE,Nipp RD,Roeland E,Ryan DP,Weekes CD,Kwak EL,Faris JE,Wo JY,Aguet F,Dey-Guha I,Hazar-Rethinam M,Dias-Santagata D,Ting DT,Zhu AX,Hong TS,Golub TR,Iafrate AJ,Adalsteinsson VA,Bardelli A,Parida L,Juric D,Getz G,Corcoran RB.Liquid versus tissue biopsy for detecting acquired resistance and tumor heterogeneity in gastrointestinal cancers.
2019; 25: 1415-1421 [PMID: 31501609 DOI: 10.1038/s41591-019-0561-9]
14 Stahler A,Heinemann V,Holch JW,von Einem JC,Westphalen CB,Heinrich K,Schlieker L,Jelas I,Alig AHS,Fischer LE,Weiss L,Modest DP,von Weikersthal LF,Decker T,Kiani A,Moehler M,Kaiser F,Kirchner T,Jung A,Stintzing S.Mutational profiles of metastatic colorectal cancer treated with FOLFIRI plus cetuximab or bevacizumab before and after secondary resection (AIO KRK 0306; FIRE-3).
2021; 149: 1935-1943 [PMID: 34310714 DOI: 10.1002/ijc.33747]
15 Woolston A,Barber LJ,Griffiths B,Pich O,Lopez-Bigas N,Matthews N,Rao S,Watkins D,Chau I,Starling N,Cunningham D,Gerlinger M.Mutational signatures impact the evolution of anti-EGFR antibody resistance in colorectal cancer.
2021; 5: 1024-1032 [PMID: 34017094 DOI: 10.1038/s41559-021-01470-8]
16 Heitzer E,Haque IS,Roberts CES,Speicher MR.Current and future perspectives of liquid biopsies in genomics-driven oncology.
2019; 20: 71-88 [PMID: 30410101 DOI: 10.1038/s41576-018-0071-5]
17 Luo W,Rao M,Qu J,Luo D.Applications of liquid biopsy in lung cancer-diagnosis,prognosis prediction,and disease monitoring.
2018; 10: 3911-3923 [PMID: 30662639]
18 Vitiello PP,De Falco V,Giunta EF,Ciardiello D,Cardone C,Vitale P,Zanaletti N,Borrelli C,Poliero L,Terminiello M,Arrichiello G,Caputo V,Famiglietti V,Mattera Iacono V,Marrone F,Di Liello A,Martini G,Napolitano S,Caraglia M,Lombardi A,Franco R,De Vita F,Morgillo F,Troiani T,Ciardiello F,Martinelli E.Clinical Practice Use of Liquid Biopsy to Identify RAS/BRAF Mutations in Patients with Metastatic Colorectal Cancer (mCRC): A Single Institution Experience.
2019; 11 [PMID: 31597339 DOI: 10.3390/cancers11101504]
19 Yi Z,Qu C,Zeng Y,Liu Z.Liquid biopsy: early and accurate diagnosis of brain tumor.
2022 [PMID: 35451698 DOI: 10.1007/s00432-022-04011-3]
20 Palacín-Aliana I,García-Romero N,Asensi-Puig A,Carrión-Navarro J,González-Rumayor V,Ayuso-Sacido á.Clinical Utility of Liquid Biopsy-Based Actionable Mutations Detected
ddPCR.
2021; 9 [PMID: 34440110 DOI: 10.3390/biomedicines9080906]
21 Vendrell JA,Mau-Them FT,Béganton B,Godreuil S,Coopman P,Solassol J.Circulating Cell Free Tumor DNA Detection as a Routine Tool forLung Cancer Patient Management.
2017; 18 [PMID: 28146051 DOI: 10.3390/ijms18020264]
22 Moreno-Manuel A,Calabuig-Fari?as S,Obrador-Hevia A,Blasco A,Fernández-Díaz A,Sirera R,Camps C,Jantus-Lewintre E.dPCR application in liquid biopsies: divide and conquer.
2021; 21: 3-15 [PMID: 33305634 DOI: 10.1080/14737159.2021.1860759]
23 Perkins G,Lu H,Garlan F,Taly V.Droplet-Based Digital PCR.In: Advances in Clinical Chemistry.Elsevier,2017: 43-91
24 Diehl F,Li M,He Y,Kinzler KW,Vogelstein B,Dressman D.BEAMing: single-molecule PCR on microparticles in waterin-oil emulsions.
2006; 3: 551-559 [PMID: 16791214 DOI: 10.1038/nmeth898]
25 Bai Y,Wang Z,Liu Z,Liang G,Gu W,Ge Q.Technical progress in circulating tumor DNA analysis using next generation sequencing.
2020; 49: 101480 [PMID: 31711827 DOI: 10.1016/j.mcp.2019.101480]
26 Thress KS,Brant R,Carr TH,Dearden S,Jenkins S,Brown H,Hammett T,Cantarini M,Barrett JC.EGFR mutation detection in ctDNA from NSCLC patient plasma: A cross-platform comparison of leading technologies to support the clinical development of AZD9291.
2015; 90: 509-515 [PMID: 26494259 DOI: 10.1016/j.lungcan.2015.10.004]
27 Diaz LA Jr,Bardelli A.Liquid biopsies: genotyping circulating tumor DNA.
2014; 32: 579-586 [PMID: 24449238 DOI: 10.1200/JCO.2012.45.2011]
28 Cabel L,Proudhon C,Mariani P,Tzanis D,Beinse G,Bieche I,Pierga JY,Bidard FC.Circulating tumor cells and circulating tumor DNA: What surgical oncologists need to know?
2017; 43: 949-962 [PMID: 28185687 DOI: 10.1016/j.ejso.2017.01.010]
29 Siravegna G,Marsoni S,Siena S,Bardelli A.Integrating liquid biopsies into the management of cancer.
2017; 14: 531-548 [PMID: 28252003 DOI: 10.1038/nrclinonc.2017.14]
30 De Rubis G,Rajeev Krishnan S,Bebawy M.Liquid Biopsies in Cancer Diagnosis,Monitoring,and Prognosis.
2019; 40: 172-186 [PMID: 30736982 DOI: 10.1016/j.tips.2019.01.006]
31 Yarden Y,Sliwkowski MX.Untangling the ErbB signalling network.
2001; 2: 127-137 [PMID: 11252954 DOI: 10.1038/35052073]
32 Sforza V,Martinelli E,Ciardiello F,Gambardella V,Napolitano S,Martini G,Della Corte C,Cardone C,Ferrara ML,Reginelli A,Liguori G,Belli G,Troiani T.Mechanisms of resistance to anti-epidermal growth factor receptor inhibitors in metastatic colorectal cancer.
2016; 22: 6345-6361 [PMID: 27605871 DOI: 10.3748/wjg.v22.i28.6345]
33 Diaz LA Jr,Williams RT,Wu J,Kinde I,Hecht JR,Berlin J,Allen B,Bozic I,Reiter JG,Nowak MA,Kinzler KW,Oliner KS,Vogelstein B.The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers.
2012; 486: 537-540 [PMID: 22722843 DOI: 10.1038/nature11219]
34 Pylayeva-Gupta Y,Grabocka E,Bar-Sagi D.RAS oncogenes: weaving a tumorigenic web.
2011; 11: 761-774 [PMID: 21993244 DOI: 10.1038/nrc3106]
35 Knickelbein K,Zhang L.Mutant KRAS as a critical determinant of the therapeutic re-sponse of colorectal cancer.
2015; 2: 4-12 [DOI: 10.1016/j.gendis.2014.10.002]
36 Lowy DR,Willumsen BM.Function and regulation of ras.
1993; 62: 851-891 [PMID: 8352603 DOI: 10.1146/annurev.bi.62.070193.004223]
37 Misale S,Yaeger R,Hobor S,Scala E,Janakiraman M,Liska D,Valtorta E,Schiavo R,Buscarino M,Siravegna G,Bencardino K,Cercek A,Chen CT,Veronese S,Zanon C,Sartore-Bianchi A,Gambacorta M,Gallicchio M,Vakiani E,Boscaro V,Medico E,Weiser M,Siena S,Di Nicolantonio F,Solit D,Bardelli A.Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer.
2012; 486: 532-536 [PMID: 22722830 DOI: 10.1038/nature11156]
38 Misale S,Di Nicolantonio F,Sartore-Bianchi A,Siena S,Bardelli A.Resistance to anti-EGFR therapy in colorectal cancer: from heterogeneity to convergent evolution.
2014; 4: 1269-1280 [PMID: 25293556 DOI: 10.1158/2159-8290.CD-14-0462]
39 Gerlinger M.Targeted drugs ramp up cancer mutability.
2019; 366: 1452-1453 [PMID: 31857471 DOI: 10.1126/science.aaz9900]
40 Russo M,Crisafulli G,Sogari A,Reilly NM,Arena S,Lamba S,Bartolini A,Amodio V,Magrì A,Novara L,Sarotto I,Nagel ZD,Piett CG,Amatu A,Sartore-Bianchi A,Siena S,Bertotti A,Trusolino L,Corigliano M,Gherardi M,Lagomarsino MC,Di Nicolantonio F,Bardelli A.Adaptive mutability of colorectal cancers in response to targeted therapies.
2019; 366: 1473-1480 [PMID: 31699882 DOI: 10.1126/science.aav4474]
41 Pietrantonio F,Vernieri C,Siravegna G,Mennitto A,Berenato R,Perrone F,Gloghini A,Tamborini E,Lonardi S,Morano F,Picciani B,Busico A,Volpi CC,Martinetti A,Battaglin F,Bossi I,Pellegrinelli A,Milione M,Cremolini C,Di Bartolomeo M,Bardelli A,de Braud F.Heterogeneity of Acquired Resistance to Anti-EGFR Monoclonal Antibodies in Patients with Metastatic Colorectal Cancer.
2017; 23: 2414-2422 [PMID: 27780856 DOI: 10.1158/1078-0432.CCR-16-1863]
42 Vidal J,Muinelo L,Dalmases A,Jones F,Edelstein D,Iglesias M,Orrillo M,Abalo A,Rodríguez C,Brozos E,Vidal Y,Candamio S,Vázquez F,Ruiz J,Guix M,Visa L,Sikri V,Albanell J,Bellosillo B,López R,Montagut C.Plasma ctDNA RAS mutation analysis for the diagnosis and treatment monitoring of metastatic colorectal cancer patients.
2017; 28: 1325-1332 [PMID: 28419195 DOI: 10.1093/annonc/mdx125]
43 Hedtke M,Pessoa Rejas R,Froelich MF,Ast V,Duda A,Mirbach L,Costina V,Martens UM,Hofheinz RD,Neumaier M,Haselmann V.Liquid profiling of circulating tumor DNA in colorectal cancer: steps needed to achieve its full clinical value as standard care.
2022; 16: 2042-2056 [PMID: 34873826 DOI: 10.1002/1878-0261.13156]
44 Morelli MP,Overman MJ,Dasari A,Kazmi SMA,Mazard T,Vilar E,Morris VK,Lee MS,Herron D,Eng C,Morris J,Kee BK,Janku F,Deaton FL,Garrett C,Maru D,Diehl F,Angenendt P,Kopetz S.Characterizing the patterns of clonal selection in circulating tumor DNA from patients with colorectal cancer refractory to anti-EGFR treatment.
2015; 26: 731-736 [PMID: 25628445 DOI: 10.1093/annonc/mdv005]
45 Strickler JH,Loree JM,Ahronian LG,Parikh AR,Niedzwiecki D,Pereira AAL,McKinney M,Korn WM,Atreya CE,Banks KC,Nagy RJ,Meric-Bernstam F,Lanman RB,Talasaz A,Tsigelny IF,Corcoran RB,Kopetz S.Genomic Landscape of Cell-Free DNA in Patients with Colorectal Cancer.
2018; 8: 164-173 [PMID: 29196463 DOI: 10.1158/2159-8290.CD-17-1009]
46 Yamada T,Matsuda A,Takahashi G,Iwai T,Takeda K,Ueda K,Kuriyama S,Koizumi M,Shinji S,Yokoyama Y,Ohta R,Yoshida H.Emerging RAS,BRAF,and EGFR mutations in cell-free DNA of metastatic colorectal patients are associated with both primary and secondary resistance to first-line anti-EGFR therapy.Int J Clin Oncol 2020; 25: 1523-1532 [PMID: 32394048 DOI: 10.1007/s10147-020-01691-0]
47 Price TJ,Peeters M,Kim TW,Li J,Cascinu S,Ruff P,Suresh AS,Thomas A,Tjulandin S,Zhang K,Murugappan S,Sidhu R.Panitumumab versus cetuximab in patients with chemotherapy-refractory wild-type KRAS exon 2 metastatic colorectal cancer (ASPECCT): a randomised,multicentre,open-label,non-inferiority phase 3 study.Lancet Oncol 2014; 15: 569-579 [PMID: 24739896 DOI: 10.1016/S1470-2045(14)70118-4]
48 Kim TW,Peeters M,Thomas A,Gibbs P,Hool K,Zhang J,Ang AL,Bach BA,Price T.Impact of Emergent Circulating Tumor DNA RAS Mutation in Panitumumab-Treated Chemoresistant Metastatic Colorectal Cancer.Clin Cancer Res 2018; 24: 5602-5609 [PMID: 29898991 DOI: 10.1158/1078-0432.CCR-17-3377]
49 Peeters M,Price T,Boedigheimer M,Kim TW,Ruff P,Gibbs P,Thomas A,Demonty G,Hool K,Ang A.Evaluation of Emergent Mutations in Circulating Cell-Free DNA and Clinical Outcomes in Patients with Metastatic Colorectal Cancer Treated with Panitumumab in the ASPECCT Study.Clin Cancer Res 2019; 25: 1216-1225 [PMID: 30487126 DOI: 10.1158/1078-0432.CCR-18-2072]
50 Sun Q,Liu Y,Liu B.Use of Liquid Biopsy in Monitoring Colorectal Cancer Progression Shows Strong Clinical Correlation.Am J Med Sci 2018; 355: 220-227 [PMID: 29549923 DOI: 10.1016/j.amjms.2017.09.009]
51 Naidoo M,Piercey O,Tie J.Circulating Tumour DNA and Colorectal Cancer: the Next Revolutionary Biomarker? Curr Oncol Rep 2021; 23: 140 [PMID: 34735665 DOI: 10.1007/s11912-021-01137-4]
52 Vlachou MS,Mauri D,Zarkavelis G,Ntellas P,Tagkas C,Gkoura S,Pentheroudakis G.Plasma ctDNA RAS status selects patients for anti-EGFR treatment rechallenge in metastatic colorectal cancer: a meta-analysis.Exp Oncol 2021; 43: 252-256 [PMID: 34591420 DOI: 10.32471/exp-oncology.2312-8852.vol-43-no-3.16592]
53 Parseghian CM,Loree JM,Morris VK,Liu X,Clifton KK,Napolitano S,Henry JT,Pereira AA,Vilar E,Johnson B,Kee B,Raghav K,Dasari A,Wu J,Garg N,Raymond VM,Banks KC,Talasaz AA,Lanman RB,Strickler JH,Hong DS,Corcoran RB,Overman MJ,Kopetz S.Anti-EGFR-resistant clones decay exponentially after progression: implications for anti-EGFR re-challenge.Ann Oncol 2019; 30: 243-249 [PMID: 30462160 DOI: 10.1093/annonc/mdy509]
54 Ciardiello D,Martini G,Famiglietti V,Napolitano S,De Falco V,Troiani T,Latiano TP,Ros J,Elez Fernandez E,Vitiello PP,Maiello E,Ciardiello F,Martinelli E.Biomarker-Guided Anti-Egfr Rechallenge Therapy in Metastatic Colorectal Cancer.Cancers (Basel) 2021; 13 [PMID: 33920531 DOI: 10.3390/cancers13081941]
55 Nakajima H,Kotani D,Bando H,Kato T,Oki E,Shinozaki E,Sunakawa Y,Yamazaki K,Yuki S,Nakamura Y,Yamanaka T,Yoshino T,Ohta T,Taniguchi H,Kagawa Y.REMARRY and PURSUIT trials: liquid biopsy-guided rechallenge with anti-epidermal growth factor receptor (EGFR) therapy with panitumumab plus irinotecan for patients with plasma RAS wild-type metastatic colorectal cancer.BMC Cancer 2021; 21: 674 [PMID: 34098908 DOI: 10.1186/s12885-021-08395-2]
56 Vangala D,Ladigan S,Liffers ST,Noseir S,Maghnouj A,G?tze TM,Verdoodt B,Klein-Scory S,Godfrey L,Zowada MK,Huerta M,Edelstein DL,de Villarreal JM,Marqués M,Kumbrink J,Jung A,Schiergens T,Werner J,Heinemann V,Stintzing S,Lindoerfer D,Mansmann U,Pohl M,Teschendorf C,Bernhardt C,Wolters H,Stern J,Usta S,Viebahn R,Admard J,Casadei N,Fr?hling S,Ball CR,Siveke JT,Glimm H,Tannapfel A,Schmiegel W,Hahn SA.Secondary resistance to anti-EGFR therapy by transcriptional reprogramming in patient-derived colorectal cancer models.Genome Med 2021; 13: 116 [PMID: 34271981 DOI: 10.1186/s13073-021-00926-7]
57 Raghav K,Morris V,Tang C,Morelli P,Amin HM,Chen K,Manyam GC,Broom B,Overman MJ,Shaw K,Meric-Bernstam F,Maru D,Menter D,Ellis LM,Eng C,Hong D,Kopetz S.MET amplification in metastatic colorectal cancer: an acquired response to EGFR inhibition,not a de novo phenomenon.Oncotarget 2016; 7: 54627-54631 [PMID: 27421137 DOI: 10.18632/oncotarget.10559]
58 Rimassa L,Bozzarelli S,Pietrantonio F,Cordio S,Lonardi S,Toppo L,Zaniboni A,Bordonaro R,Di Bartolomeo M,Tomasello G,Dadduzio V,Tronconi MC,Piombo C,Giordano L,Gloghini A,Di Tommaso L,Santoro A.Phase II Study of Tivantinib and Cetuximab in Patients With KRAS Wild-type Metastatic Colorectal Cancer With Acquired Resistance to EGFR Inhibitors and Emergence of MET Overexpression: Lesson Learned for Future Trials With EGFR/MET Dual Inhibition.Clin Colorectal Cancer 2019; 18: 125-132.e2 [PMID: 30846365 DOI: 10.1016/j.clcc.2019.02.004]
59 Xu JM,Wang Y,Wang YL,Liu T,Ni M,Li MS,Lin L,Ge FJ,Gong C,Gu JY,Jia R,Wang HF,Chen YL,Liu RR,Zhao CH,Tan ZL,Jin Y,Zhu YP,Ogino S,Qian ZR.PIK3CA Mutations Contribute to Acquired Cetuximab Resistance in Patients with Metastatic Colorectal Cancer.Clin Cancer Res 2017; 23: 4602-4616 [PMID: 28424201 DOI: 10.1158/1078-0432.CCR-16-2738]
60 Clifton K,Rich TA,Parseghian C,Raymond VM,Dasari A,Pereira AAL,Willis J,Loree JM,Bauer TM,Chae YK,Sherrill G,Fanta P,Grothey A,Hendifar A,Henry D,Mahadevan D,Nezami MA,Tan B,Wainberg ZA,Lanman R,Kopetz S,Morris V.Identification of Actionable Fusions as an Anti-EGFR Resistance Mechanism Using a Circulating Tumor DNA Assay.JCO Precis Oncol 2019; 3 [PMID: 33015522 DOI: 10.1200/PO.19.00141]
61 Hynes NE,MacDonald G.ErbB receptors and signaling pathways in cancer.Curr Opin Cell Biol 2009; 21: 177-184 [PMID: 19208461 DOI: 10.1016/j.ceb.2008.12.010]
62 Martin V,Landi L,Molinari F,Fountzilas G,Geva R,Riva A,Saletti P,De Dosso S,Spitale A,Tejpar S,Kalogeras KT,Mazzucchelli L,Frattini M,Cappuzzo F.HER2 gene copy number status may influence clinical efficacy to anti-EGFR monoclonal antibodies in metastatic colorectal cancer patients.Br J Cancer 2013; 108: 668-675 [PMID: 23348520 DOI: 10.1038/bjc.2013.4]
63 Takegawa N,Yonesaka K.HER2 as an Emerging Oncotarget for Colorectal Cancer Treatment After Failure of Anti-Epidermal Growth Factor Receptor Therapy.Clin Colorectal Cancer 2017; 16: 247-251 [PMID: 28363756 DOI: 10.1016/j.clcc.2017.03.001]
64 La Salvia A,Lopez-Gomez V,Garcia-Carbonero R.HER2-targeted therapy: an emerging strategy in advanced colorectal cancer.Expert Opin Investig Drugs 2019; 28: 29-38 [PMID: 30513002 DOI: 10.1080/13543784.2019.1555583]
65 Gharib E,Salmanipour R,Nazemalhosseini Mojarad E,Yaghoob Taleghani M,Sarlak S,Malekzade-Moghani M,Nasrabadi PN,Meiary MA,Asadzadeh Aghdaei H,Zali MR.HER2
mCRC patients with exon 20 R784G substitution mutation do not respond to the cetuximab therapy.J Cell Physiol 2019; 234: 13137-13144 [PMID: 30549033 DOI: 10.1002/jcp.27984]
66 Zhang H,Liu R,Yan C,Liu L,Tong Z,Jiang W,Yao M,Fang W,Chen Z.Advantage of Next-Generation Sequencing in Dynamic Monitoring of Circulating Tumor DNA over Droplet Digital PCR in Cetuximab Treated Colorectal Cancer Patients.
2019; 12: 426-431 [PMID: 30562681 DOI: 10.1016/j.tranon.2018.11.015]
67 Liu R,Zhao X,Guo W,Huang M,Qiu L,Zhang W,Zhang Z,Li W,Zhu X,Chen Z.Dynamic monitoring of HER2 amplification in circulating DNA of patients with metastatic colorectal cancer treated with cetuximab.
2020; 22: 928-934 [PMID: 31571151 DOI: 10.1007/s12094-019-02215-7]
68 Jacobs SA,Lee JJ,George TJ,Wade JL 3rd,Stella PJ,Wang D,Sama AR,Piette F,Pogue-Geile KL,Kim RS,Gavin PG,Lipchik C,Feng H,Wang Y,Finnigan M,Kiesel BF,Beumer JH,Wolmark N,Lucas PC,Allegra CJ,Srinivasan A.Neratinib-Plus-Cetuximab in Quadruple-WT (
) Metastatic Colorectal Cancer Resistant to Cetuximab or Panitumumab: NSABP FC-7,A Phase Ib Study.
2021; 27: 1612-1622 [PMID: 33203645 DOI: 10.1158/1078-0432.CCR-20-1831]
69 Jacobs SA,Lee JJ,George TJ,Yothers G,Kolevska T,Yost KJ,Wade JL,Buchschacher GL,Stella PJ,Shipstone A,Pogue-Geile KL,Srinivasan A,Lucas PC,Allegra CJ.NSABP FC-11: A phase II study of neratinib (N) plus trastuzumab (T) or n plus cetuximab (C) in pa-tients (pts) with ‘quadruple wild-type (WT)’ (KRAS/NRAS/BRAF/PIK3CA WT) metastatic colorectal cancer (mCRC) based on HER2 status—Amplified (amp),non-amplified (non-amp),WT,or mutated (mt).
2019; 37: TPS716-TPS716 [DOI: 10.1200/JCO.2019.37.4_suppl.TPS716]
70 IJzerman MJ,de Boer J,Azad A,Degeling K,Geoghegan J,Hewitt C,Hollande F,Lee B,To YH,Tothill RW,Wright G,Tie J,Dawson SJ.Towards Routine Implementation of Liquid Biopsies in Cancer Management: It Is Always Too Early,until Suddenly It Is Too Late.
2021; 11 [PMID: 33440749 DOI: 10.3390/diagnostics11010103]
71 Dasari A,Morris VK,Allegra CJ,Atreya C,Benson AB 3rd,Boland P,Chung K,Copur MS,Corcoran RB,Deming DA,Dwyer A,Diehn M,Eng C,George TJ,Gollub MJ,Goodwin RA,Hamilton SR,Hechtman JF,Hochster H,Hong TS,Innocenti F,Iqbal A,Jacobs SA,Kennecke HF,Lee JJ,Lieu CH,Lenz HJ,Lindwasser OW,Montagut C,Odisio B,Ou FS,Porter L,Raghav K,Schrag D,Scott AJ,Shi Q,Strickler JH,Venook A,Yaeger R,Yothers G,You YN,Zell JA,Kopetz S.ctDNA applications and integration in colorectal cancer: an NCI Colon and Rectal-Anal Task Forces whitepaper.
2020; 17: 757-770 [PMID: 32632268 DOI: 10.1038/s41571-020-0392-0]
72 Chen SH,Tsai HL,Jiang JK,Sung YC,Huang CW,Yeh YM,Chen LT,Wang JY.Emergence of RAS mutations in patients with metastatic colorectal cancer receiving cetuximab-based treatment: a study protocol.
2019; 19: 640 [PMID: 31253124 DOI: 10.1186/s12885-019-5826-7]
73 Matsuda A,Yamada T,Takahashi T,Hirata K,Nagasaka T,Ishimaru K,Sakamoto K,Koda K,Ishikawa T,Ishida H,Matsuda K,Kuramochi H,Yoshida Y,Sonoda H,Yoshida H.A Trial Protocol of Precision Medicine for Patients with RAS Wild Metastatic Colorectal Cancer Using Liquid Biopsy (RAS-liquid Study): A Prospective,Multicenter Observational Study.
2022; 6: 52-57 [PMID: 35128137 DOI: 10.23922/jarc.2021-042]
74 Ebi H,Bando H,Taniguchi H,Sunakawa Y,Okugawa Y,Hatanaka Y,Hosoda W,Kumamoto K,Nakatani K,Yamazaki K.Japanese Society of Medical Oncology Clinical Guidelines: Molecular Testing for Colorectal Cancer Treatment,4th edition.
2020; 111: 3962-3969 [PMID: 32667108 DOI: 10.1111/cas.14567]
75 Takayama Y,Suzuki K,Muto Y,Ichida K,Fukui T,Kakizawa N,Ishikawa H,Watanabe F,Hasegawa F,Saito M,Tsujinaka S,Futsuhara K,Miyakura Y,Noda H,Konishi F,Rikiyama T.Monitoring circulating tumor DNA revealed dynamic changes in
status in patients with metastatic colorectal cancer.
2018; 9: 24398-24413 [PMID: 29849949 DOI: 10.18632/oncotarget.25309]
 World Journal of Gastrointestinal Oncology2022年9期
World Journal of Gastrointestinal Oncology2022年9期
- World Journal of Gastrointestinal Oncology的其它文章
- Nutrition deprivation affects the cytotoxic effect of CD8 T cells in hepatocellular carcinoma
- Prognostic and clinicopathological value of Twist expression in esophageal cancer:A meta-analysis
- Dissecting novel mechanisms of hepatitis B virus related hepatocellular carcinoma using meta-analysis of public data
- Prediction of gastric cancer risk by a polygenic risk score of Helicobacter pylori
- Percutaneous insertion of a novel dedicated metal stent to treat malignant hilar biliary obstruction
- Construction and analysis of an ulcer risk prediction model after endoscopic submucosal dissection for early gastric cancer
